The Gene School
Empowering agricultural research
Kasetsart Genomics Workshop 2019
This workshop is now complete! Please fill out this Expression of Interest if you are interested in the next one
The course program is available here and the materials are available here.
Supported by the Australian Government (Ministry for Industry, Science and Technology) and held at the North Bangkok campus of the prestigious Kasetsart University, Empowering agricultural research through (meta)genomics will be held in Bangkok on the 25-28th June 2019.
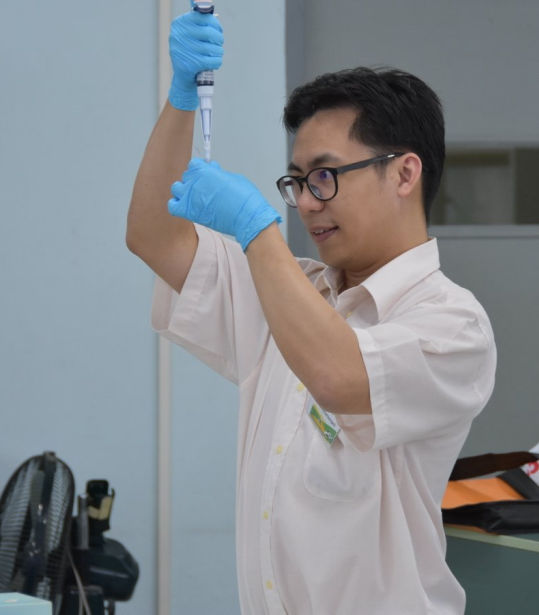
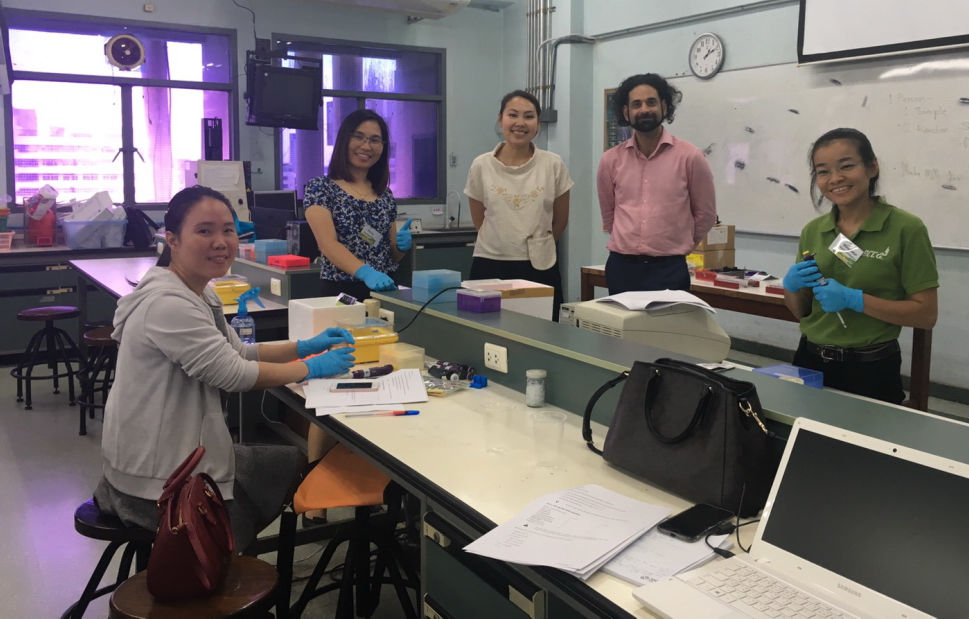
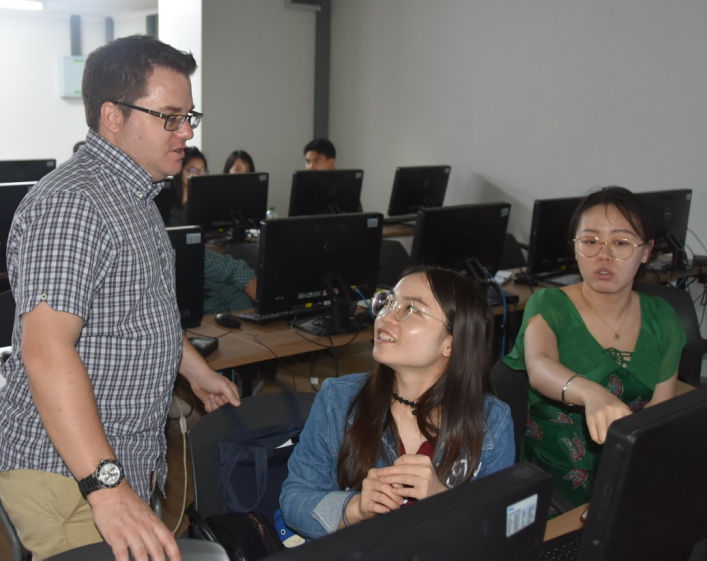
This advanced workshop follows our unique format of combining wet-laboratory techniques with scientific exploration and bioinformatics analysis. We also stream biologists and bioinformaticians so they can learn with their peers and focus on furthering their own expertise.
We start with a Research Seminar day (Prof Roger Hellens – amongst other accomplishments, of Golden Kiwi fame - providing the plenary) and early bird participants can request to submit an oral or poster presentation.
Shotgun metagenomics
Bacterial assembly and annotation
Eukaryotic genome assembly
Single assembly, annotation, and manual curation
Phylogenomics
Investigating gene families of interest
Gene expression
Differential expression and co-expression analysis
Microbiome biodiversity
Investigating biodiversity changes through 16S / ITS amplicon sequencing
Ecological diversity statistics
Identifying significant changes and what they mean
Approach
A diverse research Seminar
Our research seminar (first day of the conference) aims to inspire you but it is also a unique opportunity for selected early/mid career researchers to showcase their abilities. All early-bird registered participants can request to present
Interdisciplinary communication
Genomics is interdisciplinary. We are unlike any other course: we don't teach what you can learn by yourself. We help with your soft skills so that you can communicate effectively with peers and team members alike.
Project design
Project design is critical for genomics. During the course we use real data to show you how genomic research can solve real problems. On the last day, you show us what you've learned: Your team and you get to undertake a project design to solve a realistic problem that an industry may be facing.
Illumina and PacBio
We focus on Illumina (and PacBio), a tried and affordable technology for genomics, because we want to facilitate large scale experiments.
Galaxy and Jupyter
We use online best practice tools that implement reproducibility and notetaking: we teach you the best practices for genomics but the choice is yours!
Streaming
Biologists learn with biologists and computational scientists with computational scientists! We don't teach you to become an expert in both but an expert in
Co-Chairs of organising committee

Dr Passorn Wonnapinij
Kasetsart UniversityPassorn is a computational biologist in the faculty of Sciences. Her work focuses on mitochondrial genetics: new models of mtDNA heteroplasmy inheritance, the evolution of social amoebas, fireflies, and studying the evolution of pig through bones found in archeological sites.

Dr Alexie Papanicolaou
Hawkesbury Institute for the EnvironmentAlexie is an Assistant Professor / Senior Lecturer in Bioinformatics from the Hawkesbury Institute for the Environment (Western Sydney University); he is a computational genomicist who works (mainly) on insect genomics. He assembled and annotated multiple insect genomes (published in Nature, Molecular Ecology, Genome Biology, Nature Ecology and Evolution, and others). He led the i5k manual curation team which designed the first protocols and workflows for genome annotation and curation.
Teachers

Professor Roger Helens
Queensland University of TechnologyRoger is the Deputy Executive Director of the Institute for Future Environments of Queensland University of Technology. His career has included 14 years of working at Plant & Food Research (HortResearch) of the New Zealand government where he led a team of scientists that gave us the Golden Kiwi.

Professor Arinthip Thamchaipenet
Kasetsart UniversityArinthip is a microbial genomicist and vice-dean of the Faculty of Sciences (Kasetsart University). Her research focuses on endophytic actinomycete and their interactions with their plant hosts.

Dr Passorn Wonnapinij
Kasetsart UniversityPassorn is a computational biologist in the faculty of Sciences. Her work focuses on mitochondrial genetics: new models of mtDNA heteroplasmy inheritance, the evolution of social amoebas, fireflies, and studying the evolution of pig through bones found in archeological sites.

Dr Thomas Jeffries
Western Sydney UniversityTom is a Lecturer in Microbial Ecology at Western Sydney University with more than a decade of expertise in the genomics and biodiversity of viruses and bacteria. He co-chairs Sydney's Joint Academic Microbial Seminars and leads efforts in establishing microbial research seminars in other cities.

Prof. Aaron Darling
University of Technology, SydneyAaron is a computational biologist from the i3 research institute of the University of Technology Sydney. He has many claims to fame not least the Mauve aligner, a citation number in the 5 digit citations by the age of 35, and a scientific interest with his child's microbiome. His current work includes developing new molecular protocols for capturing microbial biodiversity, statistical bioinformatics methods to exploring the structure, function, and evolution of natural microbial communities with applications as diverse as pet foods and pig antibiotic resistance.

Dr Kay Anantanawat
University of Technology, SydneyKay is the Next Generation Sequencing facility manager for the University of Technology Sydney. After a career as a molecular entomologist, Kay is working with Aaron on creating new protocols for biodiversity assessment and running the NGS facility. Since her PhD, Kay has been extensively involved in creating and analysing NGS data. Her expertise includes creating Next Generation Sequencing libraries (RNA, DNA, and amplicon) using both commercial kits and custom methods she develops. She has hand-on experience working with sequencing instruments including 454 Roche sequencing, Illumina MiSeq and HiSeq2500.

Dr Alexie Papanicolaou
Hawkesbury Institute for the EnvironmentAlexie is an Assistant Professor / Senior Lecturer in Bioinformatics from the Hawkesbury Institute for the Environment (Western Sydney University); he is a computational genomicist who works (mainly) on insect genomics. He assembled and annotated multiple insect genomes (published in Nature, Molecular Ecology, Genome Biology, Nature Ecology and Evolution, and others). He led the i5k manual curation team which designed the first protocols and workflows for genome annotation and curation.

Associate Professor Goh Hoe Han
Universiti Kebangsaan, MalaysiaGoh is a plant molecular biologist who undertook his PhD at the University of Sheffield, UK before starting his first academic position at the Institute of Systems Biology, Universiti Kebangsaan (National University) Malaysia. Since then, he leads the Plant Functional Genomics Research Group and works on crop improvement and molecular exploration of tropical plant species using functional genomic approaches. From June 2014, he was the Head of Plant Biotechnology Centre before becoming the Head of Bioinformatics Research Centre in 2016.

Dr. Nguyen Bao Quoc
Nong Lam University, VietnamNguyen is a biotechnology researcher who completed his PhD at Kobe University, Japan in 2008 before becoming faculty at the Research Institute for Biotechnology and Environment, Nong Lam University (RIBE-NLU), Vietnam. He leads the molecular microbiology and pathogenesis group in RIBE-NLU and is the head of Human Pathogenesis Lab. His research interest is on molecular diagnostics of microbial pathogens using innovative approaches such as LAMP, RPA, realtime and PCR.

Wanwipa Vongsangnak
Kasetsart University, ThailandWanwipa completd her Ph.D. at Chalmers University of Technology, Sweden in 2009 before joining the Soochow University as an associate professor at the Center for Systems Biology in 2011. Presently, she is an associate professor at Department of Zoology, Faculty of Science, Kasetsart University, leading the Bioinformatics and Systems Biology Unit. Her research focuses on the development of new bioinformatics and systems biology approaches to study the use of microorganisms in industrial biotechnology.

Professor Federico Lauro
Nanyang Technological University, SingaporeFederico graduated from University of Padua and went on to obtain his PhD at Scripps Institute of Oceanography (SIO) at the University of California in San Diego.His driving interest is to identify how microorganisms evolve and adapt and how their functions can drive the ecological processes that are critical for sustaining the health of marine environments. His research covers both the experimental and the computational sciences – in particular, deep-sea microbiology, and the latest “omic” technologies that are essential for deriving a clear and thorough understanding of microbial communities and ecosystem function.

Dr Krithika Arumugam
Nanyang Technological University, SingaporeKrithika is a Bioinformatician at the Singapore Centre for Environmental Life Sciences Engineering (SCELSE), NTU, Singapore. With interests in metagenomics and distributed computing, she designs pipelines and analyses high throughput next generation sequencing data in high performance computing environments. Her primary research focuses on on genome recovery from metagenome assembled genomes.

Prof Mani Chellappan
Kerala Agricultural University, IndiaMani is the head of the Entomology department at the Kerala Agricultural University. As an insect physiologist, he works on the effects of nutrition and toxicity of a number of insects, in particular bees. His recent research involves understanding how the gut microbiome of invasive insects and higher vertebrates contributes to their species range and ecological adaptation.

Dr Wirulda (Nik) Pootakham
National Center for Genetic Engineering and Biotechnology, ThailandNik studied Plant Molecular Biology at Cornell University and graduated summa cum laude in 2002. She went on to pursue a PhD in Cell and Molecular Biology at Stanford University, studying a signal transduction pathway that regulates nutrient starvation responses in Chlamydomonas, a single-cell green alga. After obtaining a doctorate degree, she joined the Genomic Research Lab at the National Center for Genetic Engineering and Biotechnology (BIOTEC) and is now the Head of the Genomics Research Lab. Her work focuses on the identification of molecular markers linked to important agronomic traits for plant crop. Nik also has been involved in projects that address environmental issues of coral bleaching in the Gulf of Thailand and Andaman Sea.

Dr. Jorge Gil Angeles
University of the Philippines Los BanosJorge completed his PhD in Molecular Biology with a Minor in Bioinformatics at New Mexico State University, USA. He then worked as a post-doctoral researcher at the University of California San Francisco (UCSF) and the Northern California Institute for Research and the Education/Veterans Health Research Institute (NCIRE). Recently, he joined the Philippine Genome Center at the University of the Philippines Los Banos as the assistant leader of the functional Epigenetics and Phenomics group. His research interests include promoter research, molecular biology, epigenetics and codon usage analysis.

Prof Luca Comai
University of California at DavisLuca is professor of Plant Biology at the Genome Center of the University of California at Davis. He has B.S. equivalent from the Universita' di Bologna, Italy, and a Ph.D. in plant pathology from UC Davis. In his career, he has worked on bacterial plasmid genetics, plant biotechnology (glyphosate resistance via alteration of EPSP synthase), and genetics and genomics of polyploidy. He co-developed TILLING, a method to identify targeted mutations. Since joining UC Davis in 2006, he has focused on function and regulation of chromosomes in polyploid genomes and on stress-induced genome instability. Luca teaches the foundation genetics course at UCD using a flipped approach. He has authored over 130 publications, has an H impact factor of 72, is a Fellow of the American Association for the Advancement of Science. In 2017 he received the ASPB Innovation Prize for Agricultural Technology.

Kirk Amundson
University of California at Davis Kirk is a fourth-year PhD student in the lab of Prof. Luca Comai at the University of California, Davis. Using potato as a model system, he studies two processes that trigger genome instability in plants: first, uniparental genome elimination, a process by which sexual progeny inherit the genome of only one parent, and second, regeneration of whole plants from single cells. Although both techniques are routine in plant biology, the underlying mechanism leading to genome destabilization in both contexts remains elusive. He uses a combination of genomic and cytogenetic analyses to document the incidence and types of chromosomal restructuring events that occur in either process. By understanding how chromosomes break, his goal is to provide a framework for either preserving or engineering plant genomes.Proudly sponsored by:
This workshop was organised with funding provided by the Australian Academy of Science, on behalf of the Department of Industry, Innovation and Science. The Regional Collaborations Programme is supported by the Australian Government under the National Innovation and Science Agenda.
Thanks to our grants and also these organisations, we have been able to keep the registration affordable for early career researchers and the majority of labs.
Costs, registration, and oral presentations
Early bird registration has now expired. The new fees are 325 AUD and registrations are reviewed before being allowed on a first come, first served basis.
This covers the attendance of the hands-on workshop (Wed-Fri) and the Research Seminar day (Tuesday). All registrants can also become members of the Gene School (for free), allowing them for access to the online material and discounts to future workshops.
NB: early bird registrants will be the only ones considered for presenting in the research seminar day. This is a unique opportunity to showcase your research in front of an international audience and make connections with scientists from the region. So do book early. We will select those we think are most suitable for the event based on abstracts we request in May. If you are not selected - or don't want to give a talk - you will be encouraged to present a poster.
Registration for Thai nationals is handled by the Kasetsart University (contact us if you don't have the details), all other nationals can register either by requesting an invoice or using PayPal (no PayPal account is needed, supports credit cards, bank accounts). Please note, registrations are not confirmed until accounts are paid in full.
Registration is now full, please contact us if you are interested in the waiting listCancellations: 0 % fee (except any Paypal costs) for cancellations until the 25th of January. A 25 % fee for cancellations applies after that and before the 15th of April. No cancellations allowed after the 15th of April.
Accommodation
We recommend you find suitable accommodation and book directly. Our preferred provider is KU Home which is located 5 minutes walk from our workshop venue. Their rates are about 50 AUD/38 USD
Please note Bangkok is notorious for its traffic so we don't recommend any venues further than walking distance (also be mindful, June in Thailand is quite humid). Accommodation at KU Home and elsewhere can be booked via Booking.com, Expedia, etc.